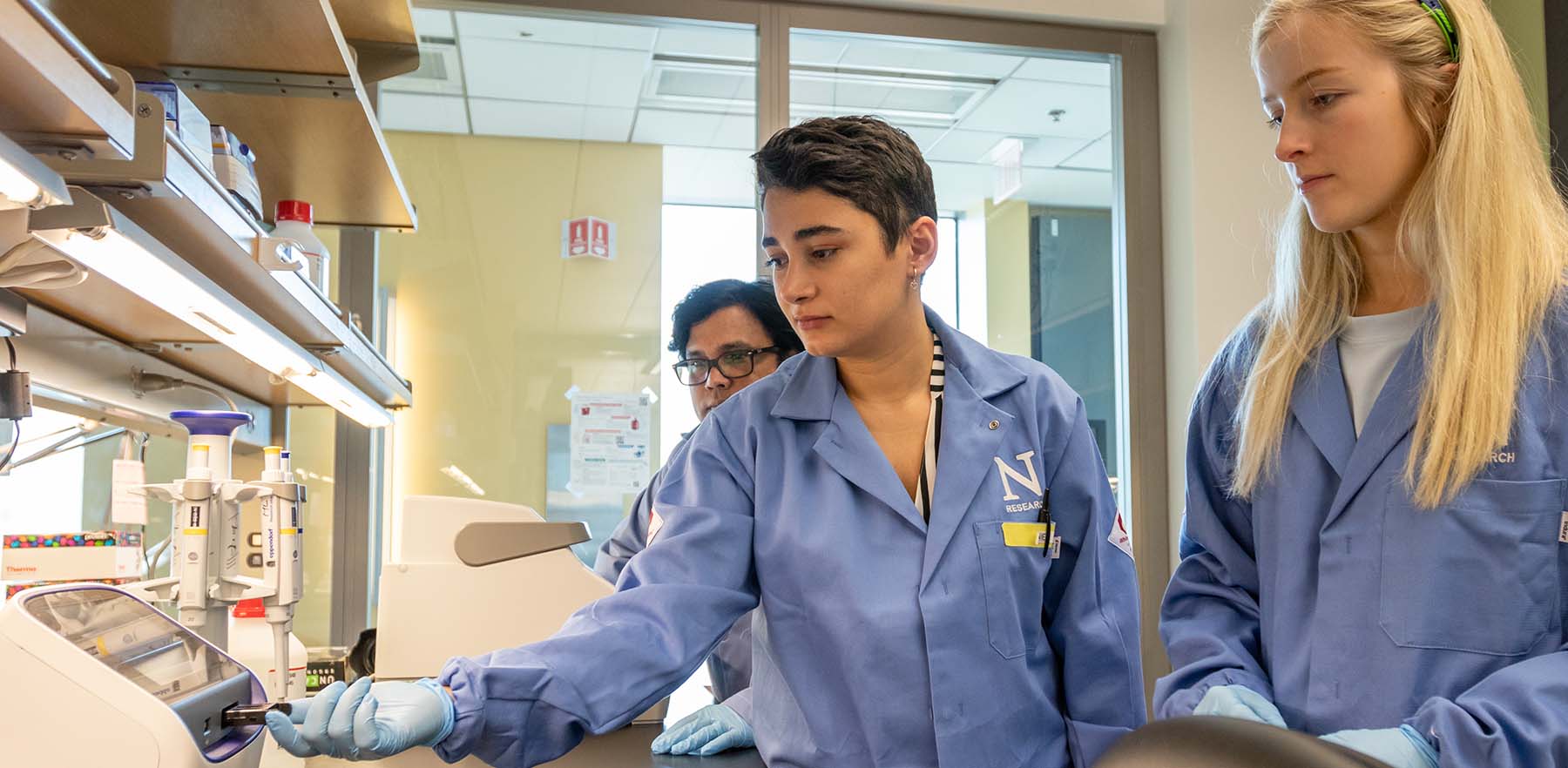



Translational Chemistry for Biomedicine
Developing new analytical technologies to combat disease
and promote wellness & health
Featured Publications
Active-reset protein sensors enable continuous in vivo monitoring of inflammation
H. Zargartalebi, S. Mirzaie, A. Ghavamin-Ejad, S.U. Ahmed, F. Esmaeili, A. Geraili, C.D. Flynn, D. Chang, J. Das, A. Abdrabou, E.H. Sargent, S. O. Kelley
Science, 2024, 386, 6726, 1146-1153.
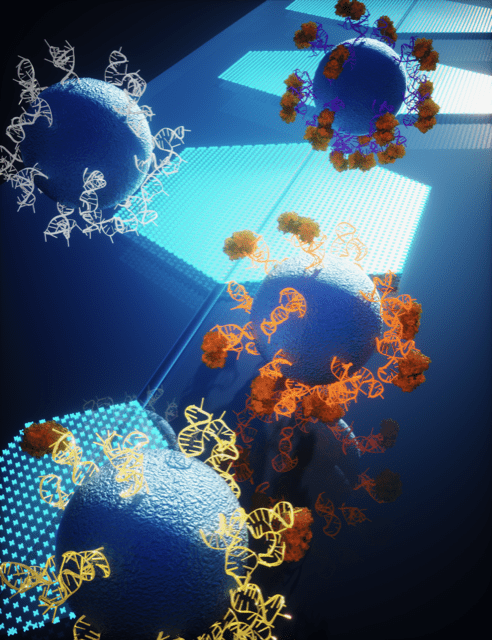
“A High-Dimensional Microfluidic Approach for Selection of Aptamers with Programmable Binding Affinities.”
D. Chang, Z. Wang, C. Flynn, A. Mahmud, M. Labib, H. Wang, A. Geraili, X. Li, J. Zhang, E. Sargent, S.O. Kelley
Nature Chemistry, 2023, 15, 773-780.
“Isolation of Tumour-Reactive Lymphocytes from Peripheral Blood via Microfluidic Immunomagnetic Cell Sorting.”
Z. Wang, S.U. Ahmed, M. Labib, H. Wang, L. Wu, F. Bavaghar-Zaeimi, N. Shokri, S. Blanco, S. Karim, K. Czarnecka-Kujawa, E.H. Sargent, A.J. McGray, M. de Perrot, S.O. Kelley
Nature Biomedical Engineering, 2023, 7, 1188-1203.
“Efficient Recovery of Potent Tumor-Infiltrating Lymphocytes Through Quantitative Immunomagnetic Cell Sorting.”
Z. Wang, S. Ahmed, M. Labib, H. Wang, X. Hu, J. Wei, Y. Yao, J. Moffat, E.H. Sargent, S.O. Kelley.
Nature Biomedical Engineering, 2022, 6, 108-117.
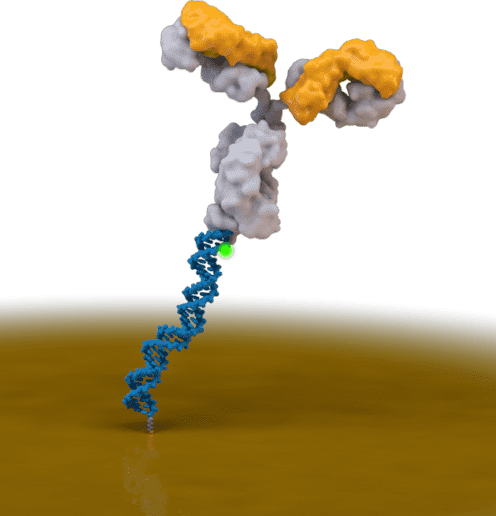
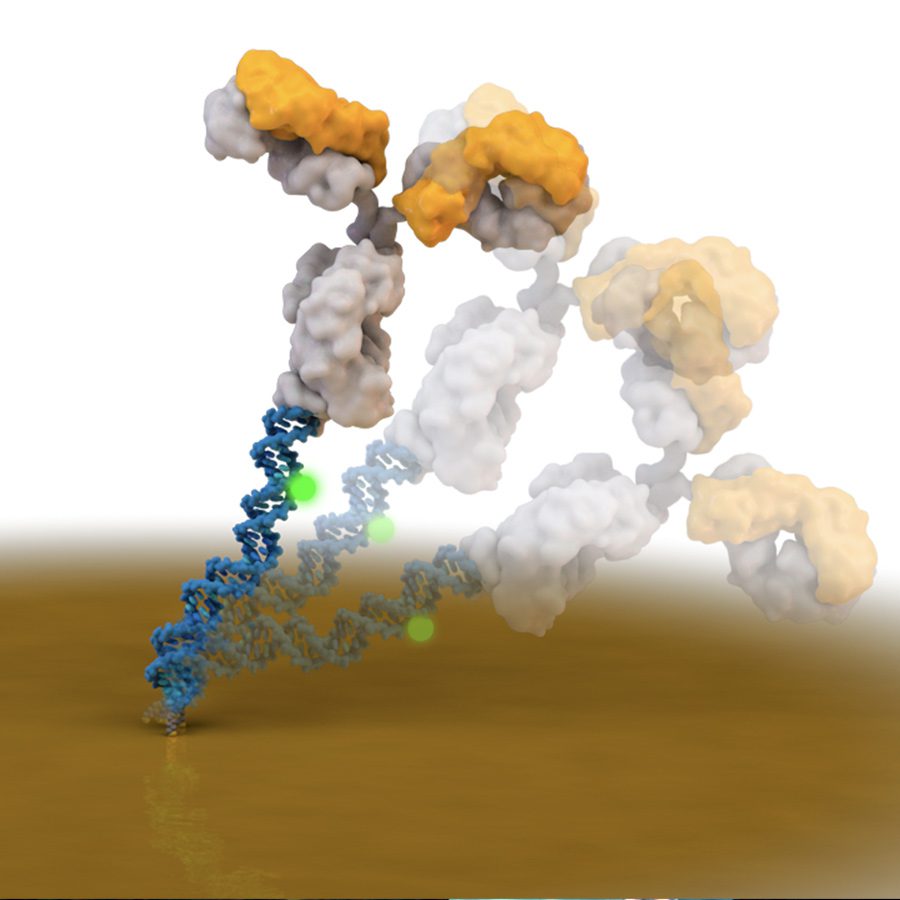
“Reagentless Biomolecular Analysis Using a Nanoscale Molecular Pendulum.”
J. Das, S. Gomis, J.B. Chen, H. Yousefi, S.Ahmed, A. Mahmud, W. Zhou, E. H. Sargent, S.O. Kelley
Nature Chemistry, 2021, 14, 428-434.
News & Press
- September 2025: Hossein Zaragartalebi presented at TEDxChicago
- September 2025: Kimberly Riordan’s paper “Dual-Chronoamperometry Drift Correction for Electrochemical Sensors” was published in ACS Sensors
- September 2025: Andrew Sedlack receives 2025 NIH National Research Service Award (NRSA) Fellowship
- September 2025: Connor Flynn’s paper “Programming Aptamer-Protein Complexation Kinetics via Modulation of G-Quadruplex Secondary Structure” was published in the Journal of the American Chemical Society
- September 2025: Xiyue (Gloria) Hu’s paper “ELOVL6 activity attenuation induces mutant KRAS degradation” was published in Nature Chemical Biology
- August 2025: Armin GhavamiNejad’s paper “Continuous insulin monitoring using and anti-body protecting zwitterionic microneedle patch” was published in Nature Biomedical Engineering
- June 2025: Zhongjie Wang’s paper “Microfluidic technologies for enhancing the potency, predictability and affordability of adoptive cell therapies” was published in Nature Biomedical Engineering
- April 2025: Connor Flynn’s paper “Nanobody Receptors Enable High-Sensitivity Monitoring of IL-6 Using Molecular Pendulum Bioanalysis” was published in Analytical Chemistry
- March 2025: Daniel Szames’ paper “Mitochondria-Targeted Temozolomide Probe for Overcoming MGMT-Mediated Resistance in Glioblastoma” was published in ChemBioChem
- January 2025: Jane Donnelley receives 2025 American Heart Association Predoc Fellowship
- December 2024: Hossein Zargartalebi’s paper “Active-reset protein sensors enable continuous in vivo monitoring of inflammation” was published in Science
Our Research
The projects underway involve aspects of diverse disciplines ranging from materials chemistry to chemical biology and nanotechnology.
Biomolecular Sensors
Advances in genomic and proteomic methods now allow classification of disease based on molecular profiling.
Rare/Single Cell Analysis
We are developing new technology platforms that allow single cell level information to be collected on billions of cells with a high level of throughput. We have applied our approaches to cell profiling to the development of assays for liquid biopsy, disease screening, cell therapy development and therapeutic target discovery.
Intracellular Molecular Delivery
Controlling the intracellular localization of synthetic molecules is essential for effective drug development. Nonetheless, rational control over intracellular trafficking of small molecules has remained a challenge. Our work has focused on mitochondrial targeting with a versatile peptide based vector that promotes efficient cellular uptake and strong mitochondrial localization.
Stay Up-to-Date with Kelley Labs
New research. New breakthroughs. New ideas.
How you keep up with the speed of science.